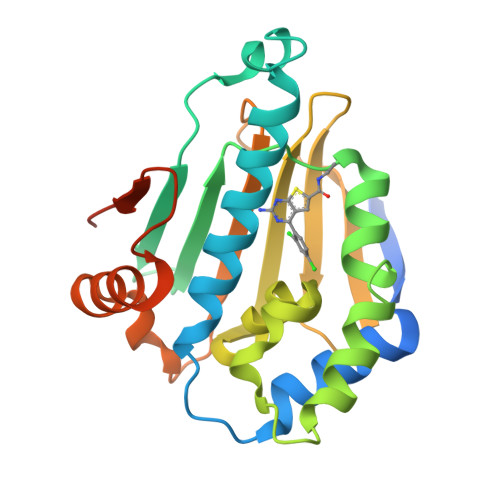
Combining Hit Identification Strategies: Fragment- Based and in Silico Approaches to Orally Active 2-Aminothieno[2,3-D]Pyrimidine Inhibitors of the Hsp90 Molecular Chaperone.
Brough, P.A., Barril, X., Borgognoni, J., Chene, P., Davies, N.G.M., Davis, B., Drysdale, M.J., Dymock, B., Eccles, S.A., Garcia-Echeverria, C., Fromont, C., Hayes, A., Hubbard, R.E., Jordan, A.M., Jensen, M.R., Massey, A., Merrett, A., Padfield, A., Parsons, R., Radimerski, T., Raynaud, F.I., Robertson, A., Roughley, S.D., Schoepfer, J., Simmonite, H., Sharp, S.Y., Surgenor, A., Valenti, M., Walls, S., Webb, P., Wood, M., Workman, P., Wright, L.M.(2009) J Med Chem 52: 4794
- PubMed: 19610616
- DOI: https://doi.org/10.1021/jm900357y
- Primary Citation of Related Structures:
2WI1, 2WI2, 2WI3, 2WI4, 2WI5, 2WI6, 2WI7 - PubMed Abstract:
Inhibitors of the Hsp90 molecular chaperone are showing considerable promise as potential molecular therapeutic agents for the treatment of cancer. Here we describe novel 2-aminothieno[2,3-d]pyrimidine ATP competitive Hsp90 inhibitors, which were designed by combining structural elements of distinct low affinity hits generated from fragment-based and in silico screening exercises in concert with structural information from X-ray protein crystallography. Examples from this series have high affinity (IC50 = 50-100 nM) for Hsp90 as measured in a fluorescence polarization (FP) competitive binding assay and are active in human cancer cell lines where they inhibit cell proliferation and exhibit a characteristic profile of depletion of oncogenic proteins and concomitant elevation of Hsp72. Several examples (34a, 34d and 34i) caused tumor growth regression at well tolerated doses when administered orally in a human BT474 human breast cancer xenograft model.
Organizational Affiliation:
Vernalis Ltd., Granta Park, Great Abington, Cambridge CB21 6GB, UK. p.brough@vernalis.com